Research Interests
Roles and mechanisms of lncRNAs in normal homeostasis and cancer development
Our research focuses on long noncoding RNAs (lncRNAs), which are transcripts longer than 500 nucleotides that lack protein-coding output. We are fascinated by lncRNAs because they outnumber protein-coding genes, yet, their functions and mechanisms of action remain poorly understood. Our primary goal is to elucidate the contribution of lncRNAs to gene regulation in normal homeostasis and during cancer development, and to understand how their localization patterns, abundance, and binding partners determine their functions.
Our previous work identified and characterized lncRNAs that mediate p53 stress response and tumor suppressive functions. We reported that the lncRNA Pvt1 (Plasmacytoma variant translocation 1), frequently altered in human cancer, harbors RNA elements that restrain the transcription of the Myc proto-oncogene in response to stress and during tumorigenesis. We also recently showed that overexpression of the lncRNA Malat1 (Metastasis-associated lung adenocarcinoma transcript 1), associated with poor prognosis in cancer patients, is an oncogenic driver of metastatic progression in a mouse model of lung cancer. With a growing appreciation that lncRNAs play central roles in normal and disease states, there is an urgent need to understand the cellular and organismal functions and to dissect the molecular mechanisms of lncRNAs. Our laboratory has extensive expertise in the development of in vitro and in vivo lncRNA-tailored genetic models and molecular tools to address these questions. We also collaborate with labs at Yale and beyond to expand our knowledge and capabilities.
1. To dissect the molecular mechanisms of transcriptional control by lncRNAs.
We recently discovered that lncRNAs modulate the expression of neighboring protein-coding genes by influencing the dynamics of their transcription. We observed that Pvt1 decreases the transcriptional output of the neighboring Myc by limiting transcriptional bursting from the Myc promoter in an RNA- and dose-dependent manner (Figure 1). To dissect the molecular mechanism, we are next interested in applying high-resolution microscopy to visualize nascent transcripts, components of the transcription machinery, and epigenetic states at the Myc/Pvt1 locus, and to perform CRISPR editing to precisely pinpoint the functional elements within Pvt1 that enables its regulatory activities.

Figure 1. Pvt1 is a chromatin-associated lncRNA that contributes to the overall local RNA concetration in the vicinity of the Myc promoter and may play a role in seeding transcriptional condensates. Myc transcriptional bursts last until the sum of Pvt1 transcripts plus newly transcribed Myc molecules reaches a threshold that triggers the RNA-mediated negative feedback mechanism that disperses transcriptional condensates and reverts the Myc locus to an inactive state. Changes in Pvt1 expression that decrease or increase the baseline local RNA concetration will alter the duration of Myc bursts, resulting in increased or decreased Myc transcriptional output and Myc levels, respectively.
Further reading:
Q. Li et al (2025) Long noncoding RNA-dependent control of Myc transcriptional bursting. Cell Reports 44, 116439.
Q. Li et al (2025) Activation of Pvt1b isoform contributes to local Pvt1 abundance to repress Myc during stress. PLoS Genet 21: e1011790.
J. Ferrer and N. Dimitrova (2024) Transcription regulation by long non-coding RNAs: mechanisms and disease relevance. Nature Reviews Molecular Cell Biology. 25(5):396-415.
C. Olivero et al (2020) p53 Activates the Long Noncoding RNA Pvt1b to Inhibit Myc and Suppress Tumorigenesis. Molecular Cell, 77(4):761-774.
2. To investigate the roles of lncRNAs in epigenetic plasticity and lineage identity.
Malat1 is an exceptional lncRNA, characterized by high conservation, abundance, and stability. Strikingly, the mechanisms that regulate Malat1 expression level, localization and function are poorly understood (Figure 2). We are interesting in investigating how Malat1 safeguards epigenetic states and lineage identity in dividing cells, and how changes in Malat1 abundance, such as its overexpression in cancer, open the door to epigenetic plasticity and lineage infidelity, which are hallmarks of cancer.

Figure 2. Malat1 abundance and localization fluctuate during the cell cucle. The mechanism and purpose of these fluctuations is not known.
3. To determine the contribution of lncRNAs to cancer initiation, metastatic progression, and drug resistance.
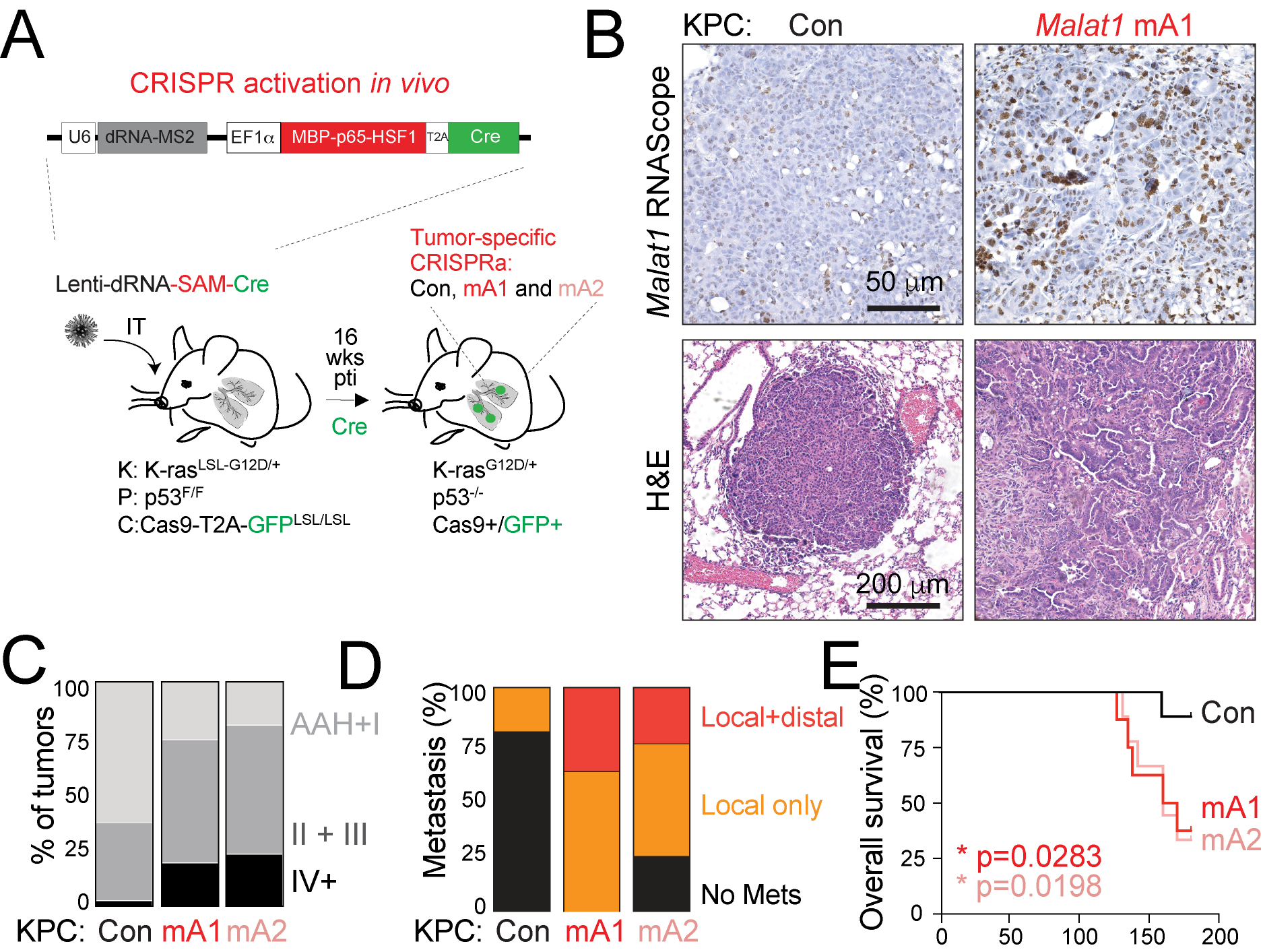
Our prior work revealed that some lncRNAs, such as Pvt1, act during early tumorigenesis to limit tumor growth, while other lncRNAs, such as Malat1, promote advanced and metastatic disease and may play a role in drug resistance (Figure 3). For trainees, interested in gaining expertise in genetically engineered mouse model of cancer, possible projects are to delve deeper into the mechanism by which Malat1 overexpression alters the tumor-immune microenvironment. Alternatively, trainees can use our powerful mouse model for in vivo CRISPRa/CRISPRi control of lncRNAs to uncover novel cancer-driving lncRNAs. These projects have the potential to highlight lncRNAs or downstream components of lncRNA pathways as promising therapeutic targets in cancer.
Figure 3. Overexpression of Malat1 in a mouse model of lung cancer by CRISPRa (A, B) leads to tumor progression (C), metastatic dissemination (D), and poor survival (E), offering a powerful tool to study humand lung adenocarcinoma, where MALAT1 is frequently elevated and highly predictive of poor prognosis.
Further reading:
E. Martinez-Terroba et al (2024) Overexpressed Malat1 drives metastasis through inflammatory reprogramming of the tumor microenvironment. Science Immunology, 9(96): eadh5462.
E. Olivero and N. Dimitrova (2020). Identification and characterization of functional long noncoding RNAs in cancer. FASEB J. 34(12):15630-15646.